MassGraph
MassGraph
A Rule-based Mass Fragment Network Database to Identify Flavonoid.
In this version (2.0), we have:
28104 flavonoids compounds which were classified as 17439 compound sets
according to the 2D-structure of each compound. It contains 7966444
"Mass Pairs" in positive ion mode and 4524415 in negative ion mode.
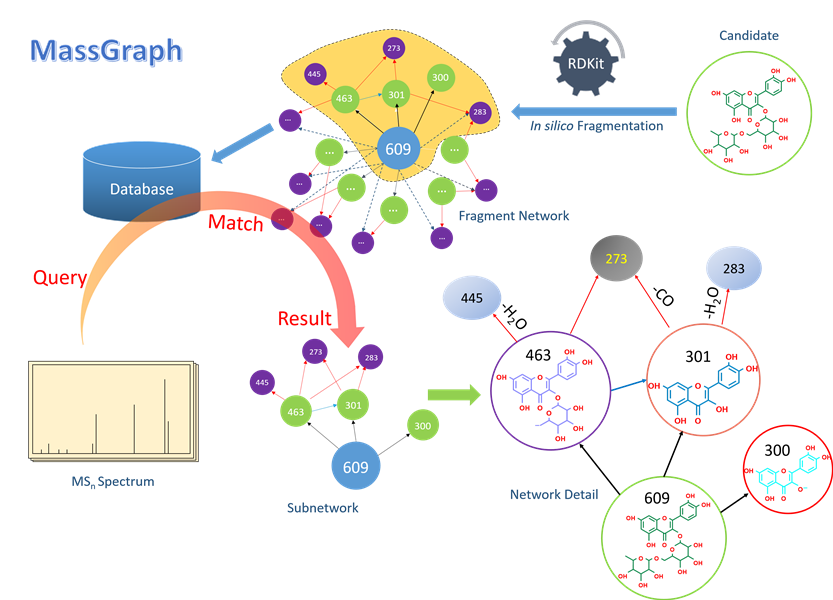
Refer to the picture below for overview of MassGraph.